Molecular Diagnostic Methods Indian Medical PG Practice Questions and MCQs
Practice Indian Medical PG questions for Molecular Diagnostic Methods. These multiple choice questions (MCQs) cover important concepts and help you prepare for your exams.
Molecular Diagnostic Methods Indian Medical PG Question 1: What is Northern blot used to detect?
- A. Protein
- B. Immunoglobulin
- C. RNA (Correct Answer)
- D. DNA
Molecular Diagnostic Methods Explanation: ***RNA***
- **Northern blot** is a laboratory technique used to detect specific **RNA** molecules among a mixture of RNA.
- It involves separating RNA fragments by **gel electrophoresis**, transferring them to a membrane, and then probing with a labeled complementary sequence.
*Protein*
- **Proteins** are typically detected using a **Western blot**, which involves similar separation and transfer techniques but uses **antibodies** as probes.
- While RNA codes for proteins, Northern blot *directly* detects RNA transcripts, not the resulting protein products.
*Immunoglobulin*
- **Immunoglobulins** (antibodies) are a type of protein, and their detection usually falls under **Western blot** or specific immunological assays like **ELISA**.
- Northern blot is specifically designed for nucleic acid analysis, not protein detection.
*DNA*
- **DNA** is detected using a **Southern blot** technique, which also involves electrophoresis, transfer to a membrane, and hybridization with a complementary probe.
- The name "Northern blot" was coined as a play on "Southern blot" because it uses similar methodology but for RNA instead of DNA.
Molecular Diagnostic Methods Indian Medical PG Question 2: What is the best investigation for identifying malaria species?
- A. Thick smear
- B. Thin smear with Giemsa (Correct Answer)
- C. QBC
- D. Thin smear with acridine orange
Molecular Diagnostic Methods Explanation: ***Thin smear with Giemsa***
- A **thin smear** allows for the visualization of **parasite morphology** within red blood cells, which is crucial for distinguishing between species of *Plasmodium*.
- **Giemsa stain** provides optimal contrast for identifying characteristic features such as **merozoites**, **trophozoites**, **schizonts**, and **gametocytes** of different malaria species.
*Thick smear*
- A **thick smear** is primarily used for **detecting the presence of malaria parasites** and for quantifying parasite density due to its higher sensitivity.
- However, because red blood cells are lysed, it **does not preserve parasite morphology** well, making species identification difficult.
*QBC*
- **Quantitative Buffy Coat (QBC) analysis** is a rapid method for detecting malaria parasites based on their fluorescence under UV light.
- While sensitive for detection, it generally **does not allow for precise species identification** due to the lack of clear morphological detail.
*Thin smear with acridine orange*
- A **thin smear stained with acridine orange** is used for rapid detection of parasites by fluorescence microscopy.
- Similar to QBC, it is **less effective for detailed morphological examination** and specific species identification compared to Giemsa-stained thin smears.
Molecular Diagnostic Methods Indian Medical PG Question 3: True about polymerase chain reaction is:
- A. Enzymatic DNA amplification (Correct Answer)
- B. Non-enzymatic DNA amplification
- C. Recombinant DNA amplification
- D. Separation of protein fragments in serum
Molecular Diagnostic Methods Explanation: ***Enzymatic DNA amplification***
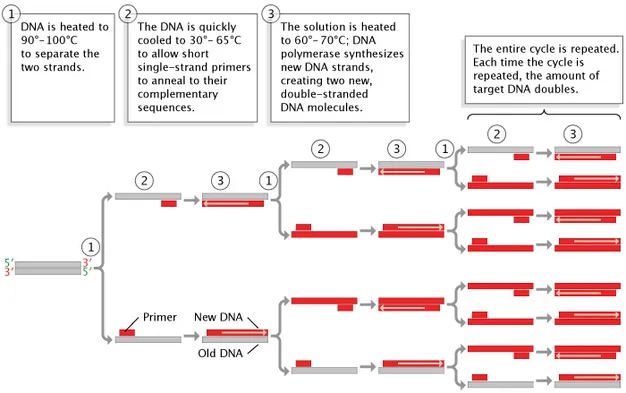
- **Polymerase Chain Reaction (PCR)** is a molecular biology technique that uses an **enzyme**, DNA polymerase (Taq polymerase), to rapidly make millions to billions of copies of a specific **DNA segment**
- The process involves cycles of denaturing the DNA (95°C), annealing primers (50-65°C), and extending the primers using **heat-stable DNA polymerase**, thus amplifying the target DNA exponentially
- Each cycle doubles the amount of target DNA, resulting in **exponential amplification**
*Non-enzymatic DNA amplification*
- PCR is fundamentally an **enzymatic process** that requires DNA polymerase enzyme
- Non-enzymatic methods of DNA amplification do not represent PCR technology
- The heat-stable **Taq polymerase** is essential for the repeated thermal cycling
*Recombinant DNA amplification*
- **Recombinant DNA technology** involves combining DNA from different sources, often for gene cloning or genetic engineering, which is distinct from PCR's primary function
- While PCR can be used to amplify recombinant DNA sequences, the technique itself is not defined as recombinant DNA amplification
- PCR simply amplifies **existing DNA sequences** without creating recombinant molecules
*Separation of protein fragments in serum*
- This describes techniques like **electrophoresis** (SDS-PAGE) or chromatography, used to separate proteins based on size or charge
- PCR deals specifically with **DNA amplification** and does not involve protein separation
- This is a completely different molecular biology technique
Molecular Diagnostic Methods Indian Medical PG Question 4: DNA sequence is determined by?
- A. Sanger sequencing (Correct Answer)
- B. PCR
- C. FISH
- D. Gel electrophoresis
Molecular Diagnostic Methods Explanation: ***Correct: Sanger sequencing***
- **Sanger sequencing** (chain-termination method) is the gold standard technique used to determine the exact order of nucleotides within a DNA molecule
- It uses dideoxynucleotides (ddNTPs) to terminate DNA strand elongation at specific bases, producing fragments of varying lengths
- These fragments are separated by capillary electrophoresis and the sequence is read based on the terminal fluorescent label
- Directly determines DNA sequence with high accuracy
*Incorrect: PCR*
- **Polymerase Chain Reaction (PCR)** amplifies specific DNA segments to create millions of copies
- It does NOT determine the sequence itself - it only makes copies of DNA
- PCR-amplified DNA can be used as a template for subsequent sequencing, but PCR itself doesn't reveal sequence information
*Incorrect: FISH*
- **Fluorescence in situ hybridization (FISH)** detects and localizes specific DNA sequences on chromosomes
- Used for chromosomal mapping and detecting chromosomal abnormalities
- Does not determine the nucleotide sequence
*Incorrect: Gel electrophoresis*
- Separates DNA fragments based on size and charge
- Used to analyze DNA but cannot determine the specific nucleotide sequence
- Useful for visualizing DNA after amplification or restriction digestion
Molecular Diagnostic Methods Indian Medical PG Question 5: Which one of the following enzymes is obtained from Thermus aquaticus bacterium that is heat stable and used in PCR at high temperature?
- A. DNA gyrase
- B. DNA polymerase III
- C. Taq polymerase (Correct Answer)
- D. Endonuclease
Molecular Diagnostic Methods Explanation: ***Taq polymerase***
- This **heat-stable DNA polymerase** is isolated from the thermophilic bacterium *Thermus aquaticus*.
- Its ability to withstand high temperatures makes it ideal for the **polymerase chain reaction (PCR)**, where DNA denaturation steps occur at elevated temperatures.
*DNA gyrase*
- **DNA gyrase** is a type II topoisomerase that introduces negative supercoils into DNA, which is important for DNA replication and transcription.
- It is not heat-stable and is not directly used for DNA amplification in PCR.
*DNA polymerase III*
- **DNA polymerase III** is the primary enzyme responsible for DNA replication in *E. coli* and other bacteria.
- It rapidly synthesizes DNA but is **not heat-stable** and would denature at the temperatures required for PCR.
*Endonuclease*
- **Endonucleases** are enzymes that cleave phosphodiester bonds within a polynucleotide chain.
- While essential for processes like DNA repair and restriction mapping, they are not primarily involved in and are not heat-stable for DNA synthesis in PCR.
Molecular Diagnostic Methods Indian Medical PG Question 6: Which of the following statements about nucleic acid amplification tests (NAATs) for STIs is FALSE?
- A. They can be used for test of cure after 3 weeks
- B. They can detect dead organisms after treatment
- C. They can be used for pharyngeal gonorrhea screening
- D. They are less sensitive than culture for rectal chlamydia (Correct Answer)
Molecular Diagnostic Methods Explanation: ***They are less sensitive than culture for rectal chlamydia***
- This statement is **FALSE**. NAATs are generally **more sensitive** than culture methods for detecting *Chlamydia trachomatis* in all anatomical sites, including the rectum.
- The high sensitivity of NAATs allows for the detection of very low bacterial loads, making them the preferred diagnostic method for many STIs.
*They can be used for test of cure after 3 weeks*
- This statement is generally **true**. While a "test of cure" (TOC) is not routinely recommended for uncomplicated *Chlamydia* or *Gonorrhea* infections due to high treatment efficacy, it can be considered in specific circumstances (e.g., persistent symptoms, pregnancy, or use of alternative regimens).
- If a TOC is performed, it should ideally be done **no sooner than 3 weeks post-treatment** to minimize potential false positives from detecting residual nucleic acids from dead organisms.
*They can detect dead organisms after treatment*
- This statement is **true**. NAATs detect the **nucleic acids (DNA or RNA)** of the target organism.
- These nucleic acids can persist in the body for a period even after the organism has been killed by treatment, leading to a positive NAAT result despite successful eradication of the infection.
*They can be used for pharyngeal gonorrhea screening*
- This statement is **true**. NAATs are the **recommended method** for detecting *Neisseria gonorrhoeae* in extragenital sites, including the pharynx.
- Pharyngeal gonorrhea is often **asymptomatic**, making screening of at-risk individuals important for public health.
Molecular Diagnostic Methods Indian Medical PG Question 7: Which of the following is a xenodiagnostic method?
- A. Intradermal test on guinea pigs for toxigenicity of Corynebacterium diphtheriae
- B. Injecting a hamster with splenic biopsy for diagnosis of leishmaniasis
- C. Injecting Aedes thorax with blood of a suspected dengue patient (Correct Answer)
- D. Rabbit ileal loop for enterotoxigenic Escherichia coli
Molecular Diagnostic Methods Explanation: ***Injecting Aedes thorax with blood of a suspected dengue patient***
- **Xenodiagnosis** involves using a live arthropod vector (such as an *Aedes* mosquito) to detect the presence of pathogens in a host by feeding the vector on the host and subsequently examining the vector for infection.
- In this method, the mosquito acts as a biological incubator or amplifier for the dengue virus, which can then be detected within the mosquito, indicating infection in the patient.
*Intradermal test on guinea pigs for toxigenicity of Corynebacterium diphtheria*
- This is an **animal pathogenicity test** to determine if the *Corynebacterium diphtheriae* strain produces toxin, but it is not xenodiagnosis as the animal is the test subject, not a vector for detection.
- The test assesses the virulence of the bacterium directly in the animal, rather than using the animal to detect an existing infection in another host.
*Injecting a hamster with splenic biopsy for diagnosis of leishmaniasis*
- This method is an example of **animal inoculation** or **culture in vivo**, where a susceptible animal is used to grow and amplify a suspected pathogen from a patient sample.
- While it uses an animal for diagnosis, it's not xenodiagnosis because the hamster is not a natural vector that feeds on the patient to acquire the pathogen.
*Rabbit ileal loop for enterotoxigenic Escherichia coli*
- This is an **animal model** used to detect the production of **enterotoxins** by *Escherichia coli*, which cause fluid accumulation in the rabbit ileum.
- This method tests for the pathogenic effect of the bacteria rather than using an arthropod vector to detect infection from a human host.
Molecular Diagnostic Methods Indian Medical PG Question 8: A frequent traveler presented with 4 days of continuous fever, abdominal pain, and bradycardia. What is the best diagnostic test to confirm the pathogen?
- A. Widal test
- B. Blood culture (Correct Answer)
- C. Urine culture
- D. Stool culture
Molecular Diagnostic Methods Explanation: ***Blood culture***
- **Blood culture** is the most sensitive and specific test for confirming **typhoid fever** in the first week of illness.
- The presence of **continuous fever** (step-ladder pattern), **abdominal pain**, and **relative bradycardia** in a traveler strongly suggests typhoid fever caused by *Salmonella Typhi*.
*Widal test*
- The **Widal test** detects antibodies against *Salmonella Typhi* antigens and is often positive later in the disease course.
- It has **limited sensitivity and specificity**, especially in endemic areas or with prior vaccination, leading to false positives and negatives.
*Urine culture*
- **Urine culture** has a low yield for *Salmonella Typhi*, as bacteria are intermittently shed in urine, usually later in the disease.
- It's primarily useful for diagnosing **urinary tract infections** or in chronic carriers of typhoid.
*Stool culture*
- **Stool culture** yield is higher in the later stages of typhoid fever, as *Salmonella Typhi* is shed in feces.
- Its sensitivity is lower than blood culture in the early acute phase when bacteremia is most prominent.
Molecular Diagnostic Methods Indian Medical PG Question 9: Which of the following statements is MOST accurate regarding herpes encephalitis?
- A. Focal neurological symptoms are common.
- B. EEG findings are nonspecific and not diagnostic.
- C. The temporal lobe is commonly involved. (Correct Answer)
- D. MRI is a key diagnostic tool.
Molecular Diagnostic Methods Explanation: ***The temporal lobe is commonly involved.***
- **Herpes simplex encephalitis (HSE)** characteristically targets the **temporal lobes** [1] and **orbitofrontal cortex**, leading to specific neurological deficits.
- This predilection for the temporal lobes often results in symptoms such as **aphasia**, **seizures**, and **memory disturbances** [1].
*Focal neurological symptoms are common.*
- While focal neurological symptoms such as **aphasia**, **hemiparesis**, and **seizures** are indeed common in HSE [1], this statement is less specific than the involvement of the temporal lobe.
- The **localization** of the infection to the temporal lobes explains why these focal symptoms are so prevalent [1].
*MRI is a key diagnostic tool.*
- **MRI findings**, particularly **T2-weighted** and **FLAIR sequences**, showing **edema** and **hemorrhage** in the temporal lobes and insular cortex, are highly suggestive of HSE.
- However, the most definitive diagnostic tool remains the detection of **HSV DNA** in the **cerebrospinal fluid (CSF)** via **PCR**.
*EEG findings are nonspecific and not diagnostic.*
- **EEG** in HSE often shows **periodic lateralizing epileptiform discharges (PLEDs)** or **focal slowing** primarily over the temporal lobes, which are highly suggestive, although not entirely diagnostic on their own.
- These findings can help guide further investigation and support a clinical diagnosis in conjunction with other tests.
Molecular Diagnostic Methods Indian Medical PG Question 10: Which of the following is not a diagnostic criterion for SIRS?
- A. Hypotension (Correct Answer)
- B. Tachypnoea
- C. Leucocytosis
- D. Tachycardia
Molecular Diagnostic Methods Explanation: ### Hypotension
- **Hypotension** is a criterion for **sepsis** and **septic shock**, but not for **SIRS** itself.
- **SIRS** criteria are based on inflammatory responses, while hypotension indicates a more severe systemic compromise.
*Tachycardia*
- **Tachycardia**, defined as a **heart rate >90 beats per minute**, is a diagnostic criterion for **SIRS** [1].
- It reflects the body's physiological stress response to a systemic inflammatory state [1].
*Tachypnoea*
- **Tachypnoea**, indicated by a **respiratory rate >20 breaths per minute** or a **PaCO2 <32 mmHg**, is a diagnostic criterion for **SIRS** [1].
- This symptom shows the body's effort to compensate for metabolic acidosis or increased oxygen demand.
*Leucocytosis*
- **Leucocytosis**, defined as a **white blood cell count >12,000/mm³** or **<4,000/mm³**, or the presence of **>10% immature neutrophils (bands)**, is a diagnostic criterion for **SIRS** [1].
- This indicates a significant systemic inflammatory response in the blood [1].
More Molecular Diagnostic Methods Indian Medical PG questions available in the OnCourse app. Practice MCQs, flashcards, and get detailed explanations.
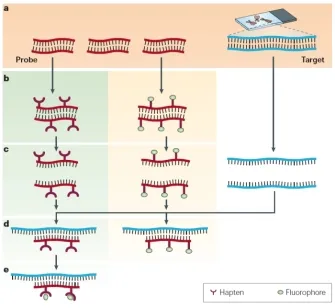 technique showing probes targeting specific DNA sequences in chromosomes)
technique showing probes targeting specific DNA sequences in chromosomes)